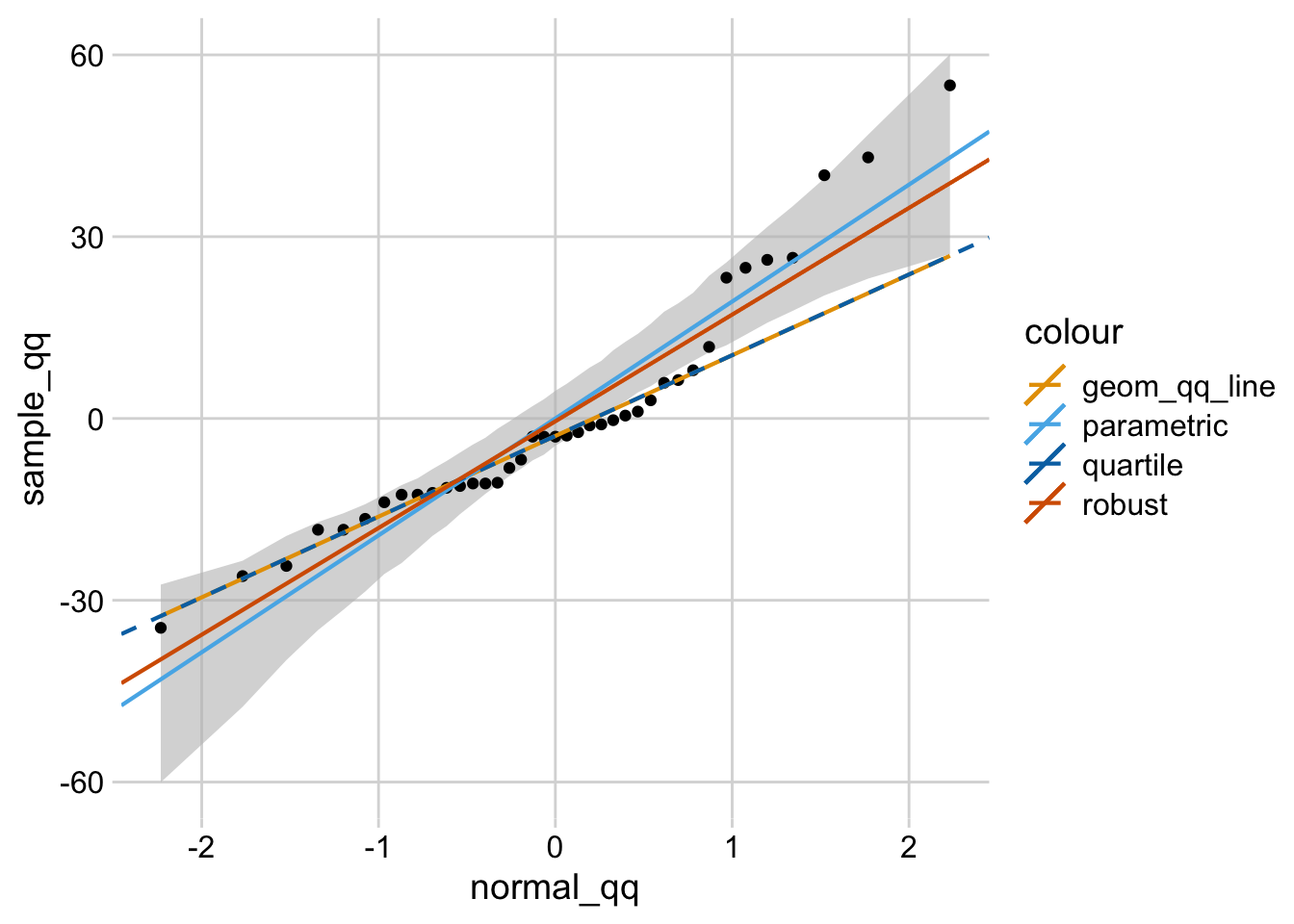
Again what to watch here is any patterns that deviate from this - particularly anything that looks curvilinear (bending at either end) or s shaped. basically what you are looking for here is the data points closely following the straight line at a 45% angle upwards (left to right). Normal Q-Q Plot: This is used to assess if your residuals are normally distributed. Or any pattern where the residuals appear non-linear (a U or upside down U shape).Īlso watch for outliers - points that are far from the general pattern of data points - as these can be influential in impacting the regression equation. What you don't want is any clear pattern where the residuals either increase/decrease in line with your predicted value. Įssentially when you look at this plot, you want the points to be in a pretty 'random' cloud pattern - ideally clustered close to the zero line (i.e. The residuals are essentially the difference between the predicted value and the actual value (i.e. Residuals - this shows the predicted value (based on the regression equation) on the X axis, and the residuals on the Y axis. here you're ideally looking for your data points to be closely clustered around a straight line (sloping up or down depending on the relationship). Predicted v Actual - this just shows how well the regression equation actually predicts the values you have in your data. Looking at the outputs it looks like you get: I don't have a lot of experience yet in StatsiQ, but do have some understanding of regression. Transform=blended_transform_factory(fig.transFigure, ax.Hi Goldie. Rc('axes.spines', top=False, right=False) from matplotlib.pyplot import subplot_mosaic, show, rcįrom ansforms import blended_transform_factoryįrom sklearn.linear_model import LinearRegression The difference between the "qq_bad" and "qq_good" plots simply has to do with selecting the column of data and passing it in as a true 1d array (instead of a 1d columnar array). The top row in the resultant figure comprises predictions & residuals for a uniform residual distribution, whereas the bottom row uses a normal distribution for errors. To highlight this, I've adapted your code sample. is a uniform distribution, so your errors aren't normal to begin with! When you're passing the residuals to qqplot it is attempting to transform each residual individually instead of an entire dataset The primary problem is that sklearn (scikit learn) expects your input to be in a 2d columnar array, whereas qqplot from statsmodels expects your data to be in a true 1d array. In my case, it does not look like the residuals follow a normal distribution, but this will change on your end because you did not set a random state when generating y_values. That is, you're looking for a histogram, not a Q-Q plot! # Import pyplot I think you were expecting to see the distribution of your errors. Regardless, if your Q-Q plot looks like a straight line, that means that your residuals and the residuals from a normal distribution match. If you look at your code, you'll see that the random terms in y_values are NOT normally distributed (you generated them with, which does not follow a normal distribution). Shouldn't the residuals be normally distributed? Hence, if the quantiles of the theoretical distribution (which is in fact normal) match those of your residuals (aka, they look like a straight line when plotted against each other), then you can conclude that the model from which you derived those residuals is a good fit. The whole idea of a Q-Q plot is to compare the quantiles of a true normal distribution against those of your residuals.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
